Background:
Food allergies are defined as a type 2 IgE-mediated immune response against certain food proteins. The balance between effector and regulatory B cells determines if an individual has an allergic or tolerant response to ingested food. This means there should be significant differences in B cell subsets and, therefore B cell regulation between healthy and allergic individuals. Therefore, an investigation into the B cell regulation in IgE production and the development of food tolerance vs food allergy is a promising trail to understand the causal pathways for food allergy. This project aims to characterize allergen-specific B cells on a transcriptomic level and investigate the differences in their B cell subsets in allergic to comparison to healthy twins.
Methods:
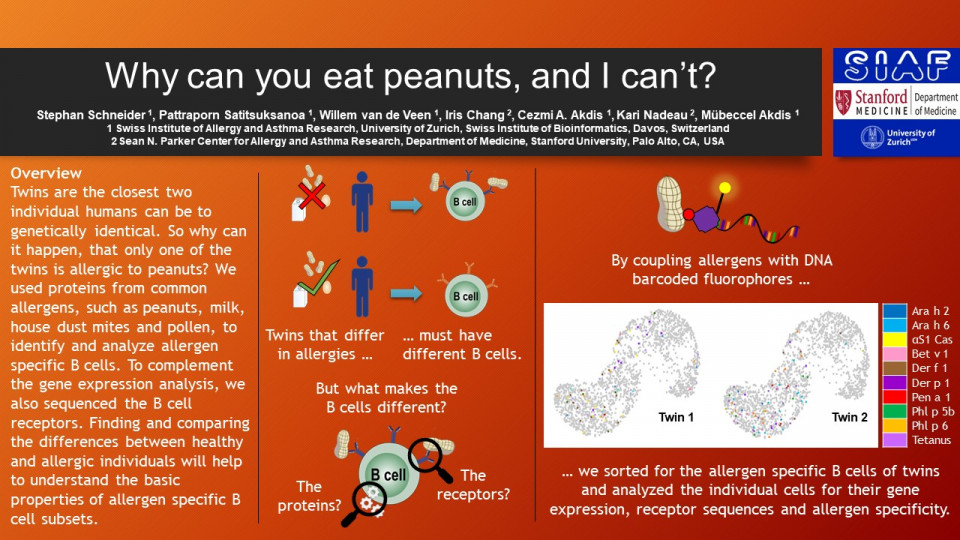
We used bio-banked PBMCs from mono- and dizygotic twins that are either food allergy concordant or discordant. The corresponding allergens were biotinylated and then coupled with a streptavidin linked to a fluorophore and an oligonucleotide barcode. This enabled us to sort for the allergen specific cells B cells and identify their allergen specificity during gene expression analysis. Additional hashtag antibodies allowed for sample multiplexing to avoid batch effects between twins. The PBMCs were sorted by FACS for cell type and allergen specificity. Using the 10x genomics Chromium Next GEM single cell sequencing technology the we generated gene expression and V(D)J libraries from the RNA of the sorted cells. These libraries were sequenced for the RNA transcriptome and also for the BCR repertoire. We analysed the sequencing data to gain insight on the abundance, clonotype and gene expression of allergen specific B cells. Comparing the cells between healthy and allergic twins will enable us to gauge the impact of V(D)J variances in allergies.